noWorkflow as an experiment management tool - Final Report
Capturing provenance into Data Science/Machine Learning workflows
This post describes our midterm work status and some achievements we have made so far in our project proposal for noWorkflow.
For a more friendly introduction to our work, please, refer to this tutorial available.
Our final code to merge is available in this repository.
Different ways of managing experiments
From our starting point at the midterm, and from our initial aspirations for the SoR, we kept on track with the goal of adding features to noWorkflow related to managing DS/ML experimental setups focusing on reproducibility.
With the emergence of IA across multiple fields in industry and academia, the subject of reproducibility has become increasingly relevant. In [1] we have an interesting description of the sources of irreproducibility in Machine Learning. All these sources are present at different stages during the project's experimental phases and may even persist in production environments, leading to the accumulation of technical debt [2]. The problem of irreproducibility is also discussed in [[3], [4]], pointing out that the velocity of deliverances usually comes at the expense of reproducibility, among other victims.
The CRISP-DM process as reviewed in [5] demonstrates that Data Science experiments follows a typical path of execution. In the same manner, [[3], [6], [7]], points out that Machine Learning pipelines are composed of well-defined layers (or stages) through its lifecycle. The emergence of IA in real world applications stressed the almost artisanal ways of creating and managing analytical experiments and reinforced that there is room to make things more efficiently.
In the search for possible approaches to the problem, we came across several projects that aimed to address these issues. Not surprisingly, multiple authors pursued the same goal, for instance [[9], [10]]. In these references, and confirmed in our survey, we found from targeted solutions to specific steps in modeling to services aiming for end-to-end AIOps management. Some are available as software packages, others as SaaS in cloud environments. In general terms, all of them end up offering features in different layers of the workflow (i.e. data, feature, scoring, and evaluation) or with different conceptualizations of reproducibility/replicability/repeatability as noticed by [11]. On one hand, this lack of standards makes any assessment difficult. On the other hand, it suggests a community in an exploratory process of a hot topic subject.
Specifically for this project, our focus is in the initial stages of computational scientific experiments. As studied in [8], in this phase, experiments are i) implemented by people as prototypes, ii) with minor focus on pipeline design and iii) in tools like Notebooks, that mix documentation, visualization and code with no required sequential structure. These three practices impact reproducibility and efficiency and are prone to create technical debts. However, tools like noWorkflow show a huge potential in such scenarios. It is promising because they i) demands a minimal setup to be functional, ii) works well with almost nonexistent workflows iii) require minimal additional intrusive code among the experimental one and iv) integrates well with Notebooks that are the typical artifact in these experiments.
According to its core team, the primary goal of noWorkflow is to "...allow scientists to benefit from provenance data analysis even when they don't use a workflow system.". Unlike other tools, "noWorkflow captures provenance from Python scripts without needing a version control system or any other environment". It is particularly interesting when we are in the scenario described above, where we lack any structured system at the beginning of experiments. In fact, after going through the docs, we can verify that noWorkflow provides:
Command-line accessibility
Seamless integration with Jupyter Notebooks
Minimal setup requirements in your environment
Elimination of the need for virtual machines or containers in its setup
Workflow-free operation
Open source license
Framework-agnostic position
Finally, in our research, we confirmed that there is an open spot in the management of scientific experiments that needs to be occupied by reproducibility. Provenance tools can help the academy and industry groups in this goal, and in this summer we focused on adding relevant features to leverage the noWorkflow in this direction.
Different tools for different needs
In our research phase, we didn't find any taxonomy that fully accommodated our review of different categories of tools providing reproducibility and experimental management. So, we could describe some tools in the following categories (freely adapted from this online references [here] and [here]):
Data and Pipeline Versioning: Platforms dealing with ingestion, processing, and exposing of features for model training and inference. They enable collaboration and discoverability of already existing Feature Sets throughout the teams and organizations. Provide provenance and lineage for data in different levels of complexity.
Metadata Stores/Experiment Trackers: They are specifically built to store metadata about ML experiments and expose it to stakeholders. They help with debugging, comparing, and collaborating on experiments. It is possible to divide them into Experiment Trackers and a Model Registry. Moreover, there are projects offering reproducibility features like hyperparameter search, experiment versioning, etc. However, they demand more robust workflows and are better suited for projects in the production/monitoring phases.
Pipeline frameworks: They operate within the realm of production, similar to Data Engineering workflows. Their usual goal is to allow any ML/AI products to be served across a wide range of architectures, and integrate all the low-hanging fruits along the way. For instance, pipelines adding hyperparameter optimization tasks, experiment tracking integrations, boilerplate containerized deployment, etc.
Deployment and Observability: They focus on deploying models for real-time inference and monitoring model quality once they are deployed in production. Their aim is to facilitate post-deployment control tasks such as monitoring feature drifts, conducting A/B testing, facilitating fast model shifts, and more.
The most remarkable aspect of this survey is that there are different tools for different phases in the life cycle of AI products. There are tools like DVC and Pachyderm that are Metadata Stores, allowing Experiment Tracking with features of tagging variables, as well as Data and Pipeline tracking. They are the most similar tools to noWorkflow in functionality. However, DVC possesses a more complex framework in dealing with different 'types' of tags, and relies on command line tools to extract and analyze tagged variables. Also, it depends strongly on git and replicate the git logics. Pachyderm requires a more sophisticated setup at the start, relying on containers and a server. It is an obstacle to small and lean prototypes, requiring installation of a docker image, and all friction on managing it.
There are other tools, like MLFlow and Neptune that pose themselves as Model Experiment Versioning with features of Monitoring and Deployment. They also have elements of pipeline frameworks, offering full integration and boiler plates for seamless integration with cloud platforms.
Pipelines are a vast field. They are AWS SageMaker, Google Vertex, DataRobot and Weights & Biases, among others. All of them offer features helping in all categories, with a strong focus on exploring all automation that can be offered to the final user, suggesting automatic parameter tuning, model selection, retraining, data lineage, metadata storing, etc.
Finally, Deployment and Observability frameworks are in the deployment realm, which is another stage far removed from prototypical phases of experiments. They come into the scene when all experimental and inferential processes are done, and there is an AI artifact that needs to be deployed and monitored. Such tools like Seldon, H2O, Datarobot do this job, again, with some features of Hyperparameter tuning, pipeline frameworks, data and pipeline tracking.
In light of this, when considering management and operation of experiments, we have a reduced sample of alternatives. Among them, Notebook integration/management are rare. Some of them rely on other tools like Git or enforces an overhead in the coding/setup with reserved keywords, tags and managerial workflows that hinder the process.
At first sight, our "informal" taxonomy positions noWorkflow as a Data/Pipeline Versioning and Metadata Store/Experiment Tracker. It is not a Pipeline Framework which works like a building block, facilitating the integration of artifacts at production stages. It is not a Deployment and Observability framework, because they are in the post-deployment realm, which is another stage far removed from prototypical phases of experiments.
Desiderata
As mentioned earlier, a typical workflow in DS/ML projects is well described by the CRISP-DM [5] and precede phases of deployment and production in the whole lifecycle of DS/ML projects.
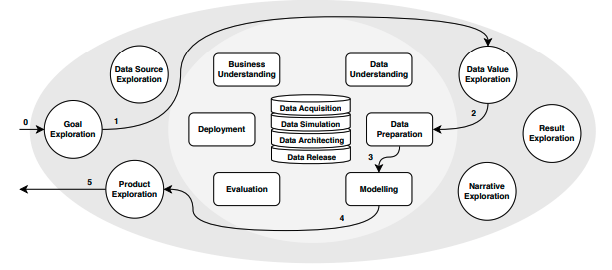
Fig 1: CRISP-DM example of trajectory through a data science project
Briefly speaking, a workflow starts when a user creates a Jupyter Notebook and starts writing code. Usually, he/she imports or selects data from a source, explore features which are expected to have the highest inference potential, tunes some parameters to set up its training, trains and evaluates the predictive power of the model through different metrics. At this final step, we have delineated a trial. This trial result can suggest further improvements and new hypotheses about data, features, model types and hyperparameters. Then, we have a new experiment in mind that will result in a new trial.
When this process repeats multiple times, a researcher may end with different notebooks storing, each one, a different experiment. Each notebook has multiple hyperparameters, modeling choices and modeling hypotheses. Otherwise, the experimenter may have a unique notebook where different experiments were executed, in a nonlinear order between the cells. This former case is pointed out in [8], where Notebook flexibility makes it difficult to understand which execution order resulted in a specific output.
In a dream space, any researcher/team would have benefited at most if they could
a) in a running Notebook, being able to retrieve all the operations that contributed to the result of a variable of interest. In this case, modifications applied in the inputs or in the order of operations would be easily detectable. In the same way, any nonlinear execution that interferes in a control result.
b) Compare trials after different experiments. After experimenting with different hypotheses about hyperparameters, features or operation order, the user should easily compare the history of two trials and spot differences.
c) Retrieve a target variable among different trials that were executed in the context of an experiment. After proceeding with multiple experimental trials, users should be able to compare the results that are stored in different Notebooks (or even not).
d) Be as much "no workflow" as possible. All the former requisites should be possible with minimal code intervention, tags, reserved words or any active coding effort.
With these goals in mind, we worked on our deliverables and used the experiment carried out by [12] as a guideline to validate the new noWorkflow features.
Deliverables
In this session, we will describe what we have implemented during this summer.
We started on tagging cells and variables and then navigating through its pre-dependencies, or all other variables and function calls that contributed to its final value. This was a fundamental step that allowed us to evolve to create features that are really useful in day-to-day practice.
From the features of tagging a cell and tagging a variable, we evolved to the following features (an interactive notebook is available here):
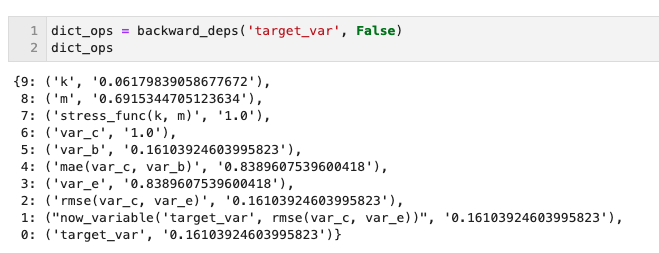
- backwards_deps('var_name', glanularity_level) : returns a dictionary storing operations/functions calls and their associated values that contributed to the final value of the tagged variable. Glanularity_level allows to set if the internal operations of the functions must be included or not.
global_backwards_deps('var_name', glanularity_level) : does the same as backwards_deps, but from all different tagging and re-tagging events in the notebook. It allows to retrieval of the complete operation of a tagged variable across all executed cells in the notebook
store_operations(trial_id, dictionary_ops) : save the current trial in order to make further comparisons with other experiments. The dictionaries aren't stored in the .noworkflow/db.sqlite, but in a shelve object named *ops.db* in the current notebook local folder.
resume_trials() : to support the management of experiments, the user can see the trial_ids of all experiments stored in the ops.db available for comparison/analysis.
trial_intersection_diff(trial_id1, trial_id2) : all mutual variables/funcion_calls between two experiments have its scalar values compared
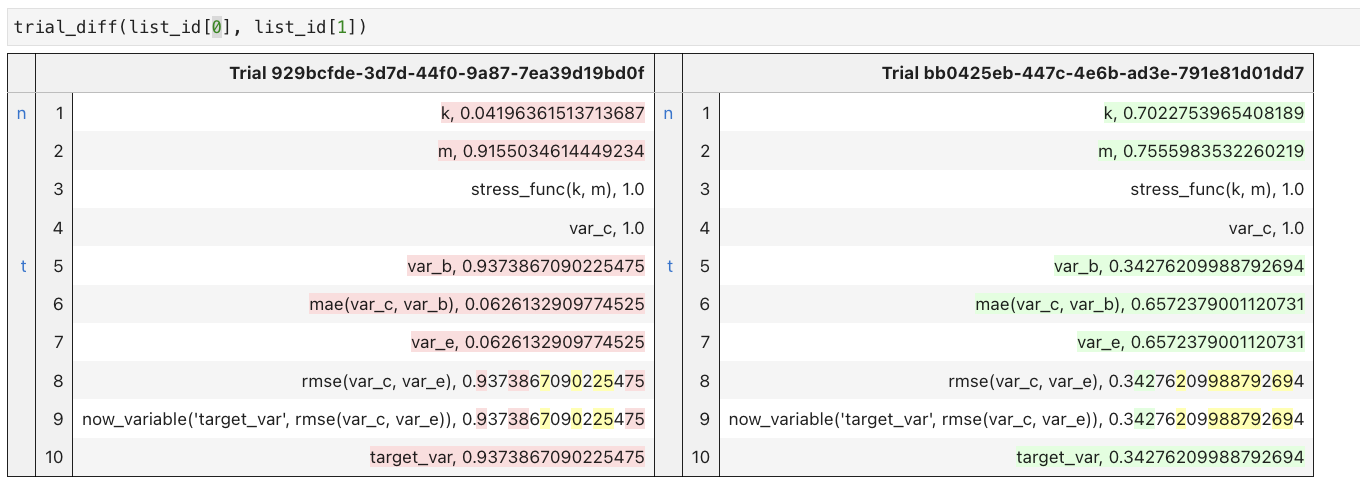
- trial_diff(trial_id1, trial_id2) : The values of variables and function calls are exhibited in a diff file format, emphasizing the operations' order. The goal here is to show that between the two experiments, the order of operations was different. Again, only scalar values are exhibited. More complex data structures (matrices, vectors, tensors, etc.) are only signaled as 'complex_type'
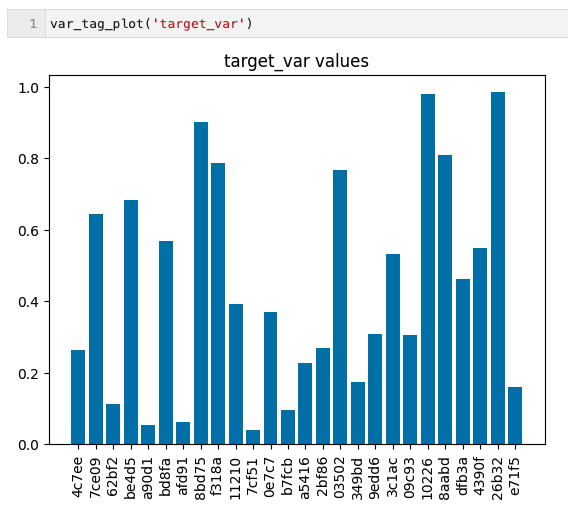
- var_tag_plot('var_name') : Chart the evolution of a given variable across multiple trials in the database. In this case, all experiments stored in ops.db and tagged as *target_var* have their values plotted
- var_tag_values('var_name') : Provides access to pandas.dataframe var_name entries with correspondent values across different trials.

Challenges
As expected, we had unexpected findings along the project. Bellow, we delve into the most significant challenges we had to face:
Jupyter notebooks allow a nonlinear execution of small parts of code through cells. More than once, we had to align about how to create functionalities to attend different scenarios that were unexpected. One example was the backwards_deps() and global_backwards_deps() functions. The latter function was born to cover the case where the user wants all dependencies rather than the local cell dependencies.
Despite the high quality of the current version of the package, the project needs documentation, which slows down the analysis of any new development. In this project, the aid of mentors was crucial at some points where a deeper knowledge was needed.
What is the vocation of noWorkflow? At some points in the project, we had to discuss forcing some kind of workflow over the user. And it would go against the philosophy of the project.
When working on comparing results, especially in DS/ML fields, complex types arise. Numerical vectors, matrices, and tensors from NumPy and other frameworks, as well as data frames, can't be properly manipulated based on our current approach.
The dilemma of focusing on graphic visual features versus more sophisticated APIs. More than once, we needed to choose between making a visual add-on to Jupyter or implementing a more complete API.
The current version of Jupyter support in noWorkflow doesn’t integrate well with Jupyter Lab. Also, even the IPython version has new versions, and noWorkflow needs to adapt to a new version.
Future Improvements
Given our current achievements and the insights gained along the project, we would highlight the following points as crucial future roadmap improvements:
Add a complex type treatment for comparisons. Today, visualizing and navigating through matrices, data frames, tensors, isn't possible with noWorkflow, although the user can do by its own means.
Integrate the dictionaries storing sequences of operations from shelve objects to a more efficient way of storage and retrieval.
Make it easier for users to manage (store, retrieve, and navigate) through different trials.
Add graphical management instead of relying upon API calls only.
Evolve the feature of tagging cells.
When tagging a model, save its binary representation to be recovered in the future.
Adding the capability of tracking the local dataset reading. Currently, it is possible to track changes in the name/path of the dataset. However, any modification in the integrity of a dataset is not traceable.
What I've learned
This was a great summer with two personal discoveries. The first one was my first formal contact with the Reproducibility subject. The second was to fully contribute with an Open Source project. In the research phase, I could get in touch with the state-of-the-art of reproducibility research and some of it nuances. In the Open Source contributing experience, I could be mentored by the core team of the noWorkflow and exercise all the skills required in doing high level software product.
Acknowledgments
I would like to thank the organization of Summer of Reproducibility for aiding this wonderful opportunity for interested people to engage with Open Source software. Also, thanks to the core team of noWorkflow for supporting me in doing this work.
Bibliography
[1] [O. E. Gundersen, K. Coakley, C. Kirkpatrick, and Y. Gil, “Sources of irreproducibility in machine learning: A review,” arXiv preprint arXiv:2204. 07610.]
[2] [D. Sculley et al., “Machine Learning: The High Interest Credit Card of Technical Debt,” in SE4ML: Software Engineering for Machine Learning (NIPS 2014 Workshop), 2014.]
[3] [P. Sugimura and F. Hartl, “Building a reproducible machine learning pipeline,” arXiv preprint arXiv:1810. 04570, 2018.]
[4] [D. Sculley et al., “Hidden technical debt in machine learning systems,” Adv. Neural Inf. Process. Syst., vol. 28, 2015.]
[5] [F. Martínez-Plumed et al., “CRISP-DM twenty years later: From data mining processes to data science trajectories,” IEEE Trans. Knowl. Data Eng., vol. 33, no. 8, pp. 3048–3061, 2019.]
[6] [N. A. Lynnerup, L. Nolling, R. Hasle, and J. Hallam, “A Survey on Reproducibility by Evaluating Deep Reinforcement Learning Algorithms on Real-World Robots,” in Proceedings of the Conference on Robot Learning, L. P. Kaelbling, D. Kragic, and K. Sugiura, Eds., in Proceedings of Machine Learning Research, vol. 100. PMLR, 30 Oct--01 Nov 2020, pp. 466–489.]
[7] [A. Masood, A. Hashmi, A. Masood, and A. Hashmi, “AIOps: predictive analytics & machine learning in operations,” Cognitive Computing Recipes: Artificial Intelligence Solutions Using Microsoft Cognitive Services and TensorFlow, pp. 359–382, 2019.]
[8] [J. F. Pimentel, L. Murta, V. Braganholo, and J. Freire, “Understanding and improving the quality and reproducibility of Jupyter notebooks,” Empirical Software Engineering, vol. 26, no. 4, p. 65, 2021.]
[9] [D. Kreuzberger, N. Kühl, and S. Hirschl, “Machine Learning Operations (MLOps): Overview, Definition, and Architecture,” IEEE Access, vol. 11, pp. 31866–31879, 2023.]
[10] [N. Hewage and D. Meedeniya, “Machine learning operations: A survey on MLOps tool support,” arXiv preprint arXiv:2202. 10169, 2022.]
[11] [H. E. Plesser, “Reproducibility vs. replicability: a brief history of a confused terminology,” Front. Neuroinform., vol. 11, p. 76, 2018.]
[12] [Z. Salekshahrezaee, J. L. Leevy, and T. M. Khoshgoftaar, “The effect of feature extraction and data sampling on credit card fraud detection,” Journal of Big Data, vol. 10, no. 1, pp. 1–17, 2023.]